Local Analysis of mAb Sequences (LAMBS)
Developing monoclonal antibody therapeutics always involves assessing candidate sequences. There are generally standard ways of doing this, but most require either uploading your proprietary sequences to some online server (which you should never do), re-inventing the wheel by implementing your own analysis, or paying for expensive software.
Local analysis of mAb sequences, LAMBS, is different: it’s free, private, open source, and doesn’t require installation, setup, or bioinformatics expertise.
It sounds too good to be true, but all of this is possible with a single HTML file you download and run securely in a web browser on your own computer. Embedded JavaScript and reference data power the analysis, while simple CSS and HTML elements orchestrate the layout and interactivity.
LAMBS is under active development and currently features:
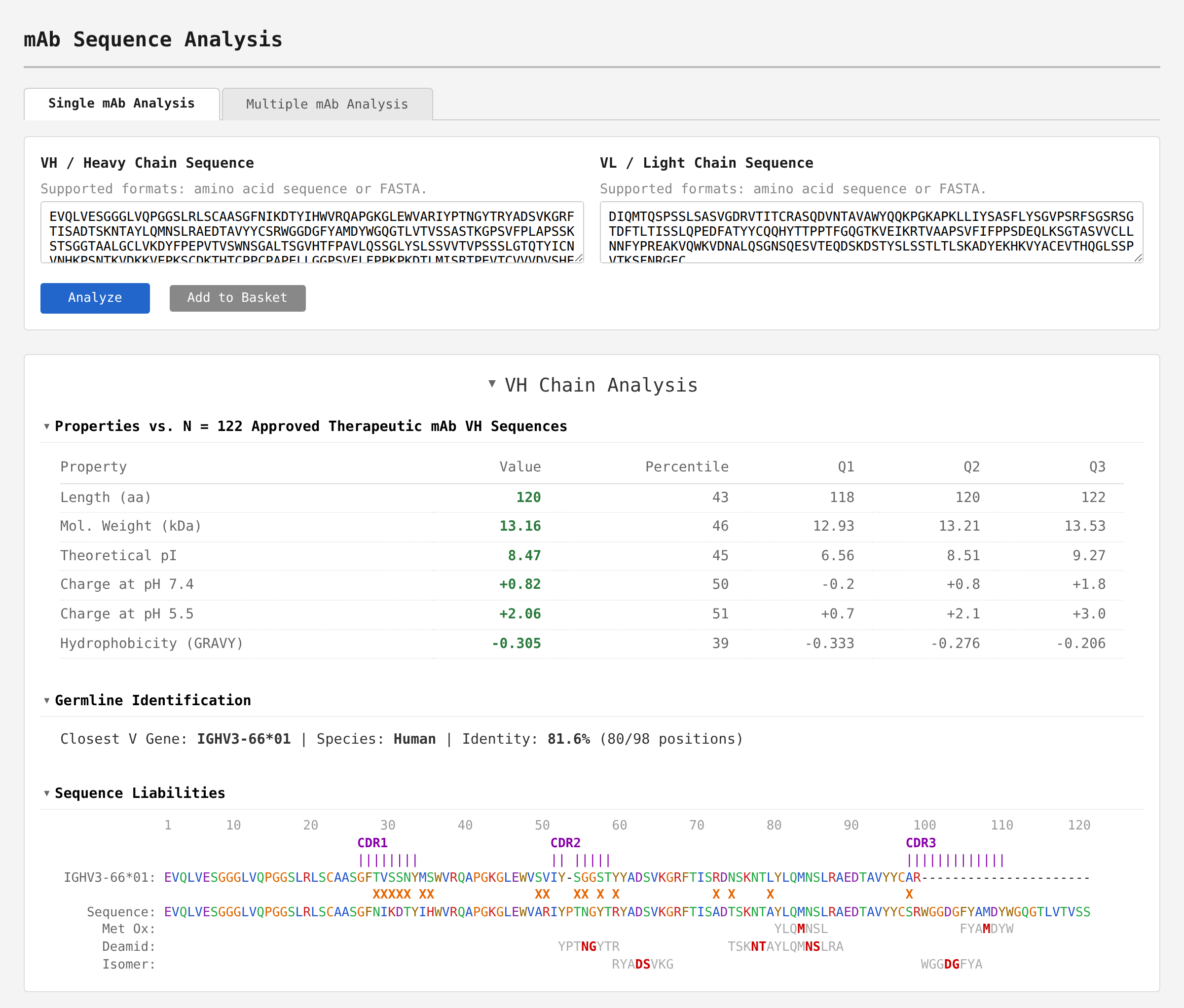
- VH and VL chain property analysis with comparisons to approved therapeutic mAbs.
- Germline gene and constant region annotation and alignments.
- Developability liability identification e.g. deamidation, isomerization, glycosylation, unpaired cysteines, and more.
- Batch analysis with sequence clustering, multiple sequence alignment, AA conservation plots, and consensus sequences.
- Privacy: everything runs locally in the browser without tracking or data being sent anywhere.
Ready to try it out? Download the index.html file from the latest release and open it in your web browser. That’s it. And don’t worry if you don’t have sequences on hand, it comes with examples preloaded for you to explore.
Interested in contributing to the project? Feel free to open an issue, submit a PR, or contact me.
What do you think? Did I miss anything? Contact me or discuss on Twitter!
This work by
Derek
Croote
is licensed under
CC BY-NC 4.0


